Core Facilities
 The Membrane Protein Structural Dynamics Consortium (MPSDC) core facilities are designed to feed and interconnect with the individual projects in a highly interactive way. They are aimed at elucidating fundamental issues about membrane protein systems by providing service and expertise in five critical areas: membrane protein expression, the establishment of chemical synthesis capabilities, the generation of a large variety of crystallization chaperones and other target binders, the consistent fulfillment of computational and modeling needs, and the generation of new tools in the phage display synthetic toxin pipeline.
The Membrane Protein Structural Dynamics Consortium (MPSDC) core facilities are designed to feed and interconnect with the individual projects in a highly interactive way. They are aimed at elucidating fundamental issues about membrane protein systems by providing service and expertise in five critical areas: membrane protein expression, the establishment of chemical synthesis capabilities, the generation of a large variety of crystallization chaperones and other target binders, the consistent fulfillment of computational and modeling needs, and the generation of new tools in the phage display synthetic toxin pipeline.
Computational Modeling
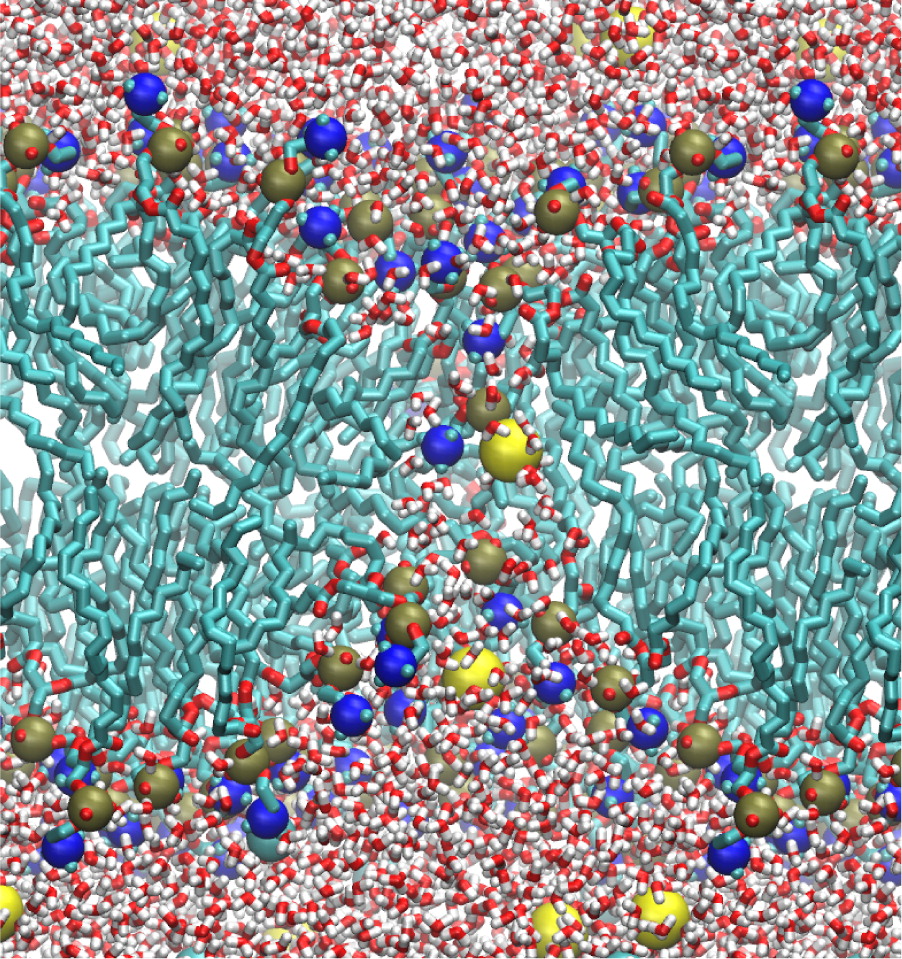

One of the key concepts in the design of the present Consortium is out ability to rationalize through computational means a vast amount of static and dynamics structural data with functional information. The computational Core is designed to generate, implement, validate and distribute state-of-the-art “tools” required for the studies of complex membrane protein systems. This includes expertise and services of well-established tools, novel methods and technologies that will support the experimental studies, such as methods for analysis of spectroscopic data, and ground work on basic computational staples like force fields and biomolecular simulations. All the computational methodologies will be readily made available to the broader scientific community through our website.
Spectroscopy and Instrumentation
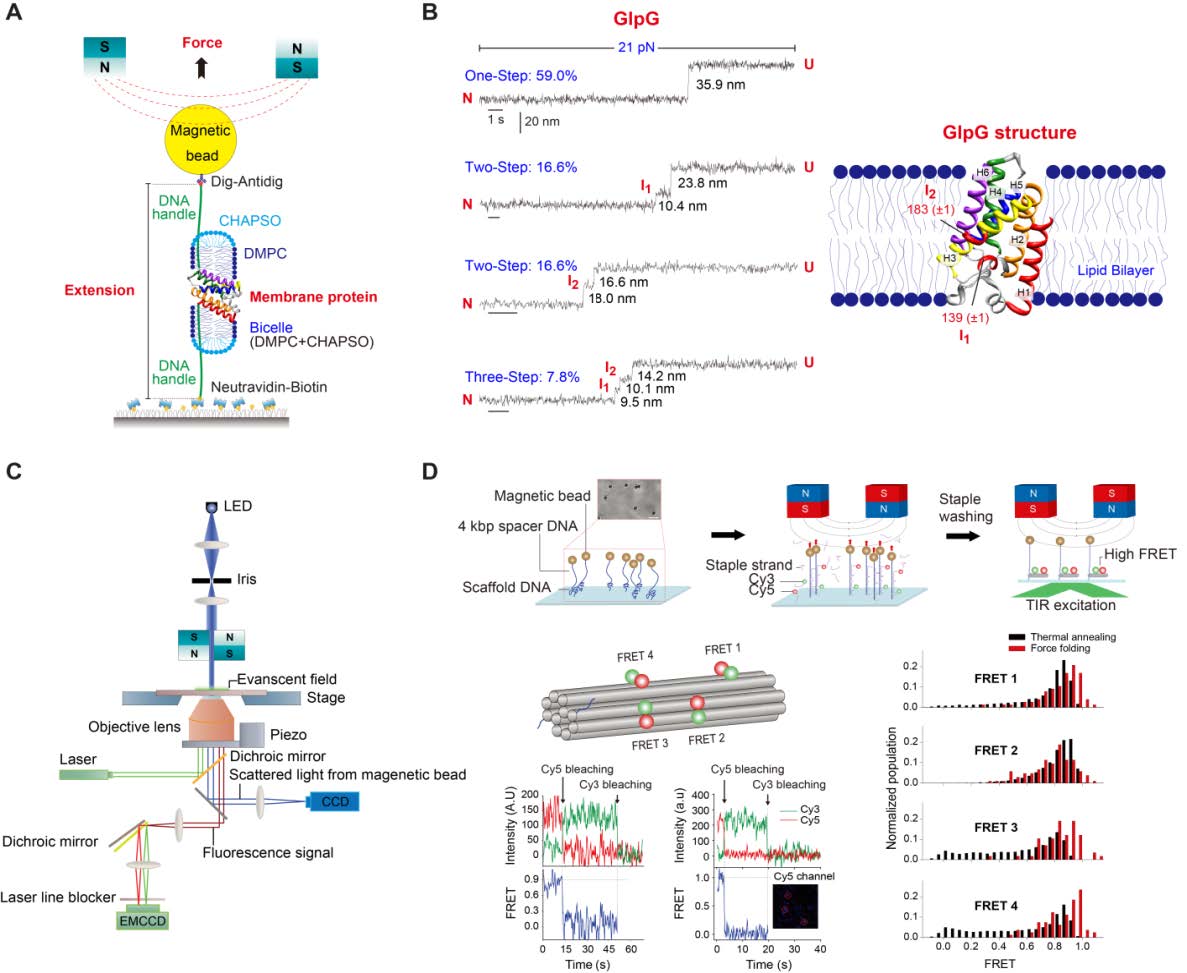
The main objective of the Membrane Protein Structural Dynamics consortium is to obtain a mechanistic understanding of how the structural dynamics of membrane proteins generate its function. The experimental study of structural dynamics is approached with a variety of spectroscopic methods and biophysical procedures that are in constant development and improvement. The objective of the Instrumentation Spectroscopy Core (SIC) is two-fold: first, to improve presently available instruments, develop new instruments, or methods and procedures aimed at improving on signal to noise ratio, specificity and time resolution of spectroscopic techniques and combining them with functional techniques, such as electrophysiology. The second objective is to provide service to the MPSD members, and the community at large, with the new developments as well as with the spectroscopic and functional techniques presently installed.
Membrane Protein Production and Chemistry

The Protein Production and Chemistry Core (PCC) does the heavy lifting for the MPSDC in support of the experimental aspects. It goal is to optimize protein expression in order to provide appropriate membrane proteins and novel spectroscopic probes for the investigators of the Consortium, and reagents, tools and protocols to the community. We consider every project a collaboration and work intimately with the investigators to develop optimized expression systems. While bacterial and cell-free expression systems are the best for high yield expression, we use eukaryotic systems, yeast, insect and mammalian cells, to generate the proteins when they provide the best approach for achieving the properly folded protein. We are also developing new spectroscopic probes as unnatural amino acids (UAAs), which are well suited for incorporation in single cells or for single molecule approaches.
Synthetic Antigen Binder (SAB) Generation and Crystallography
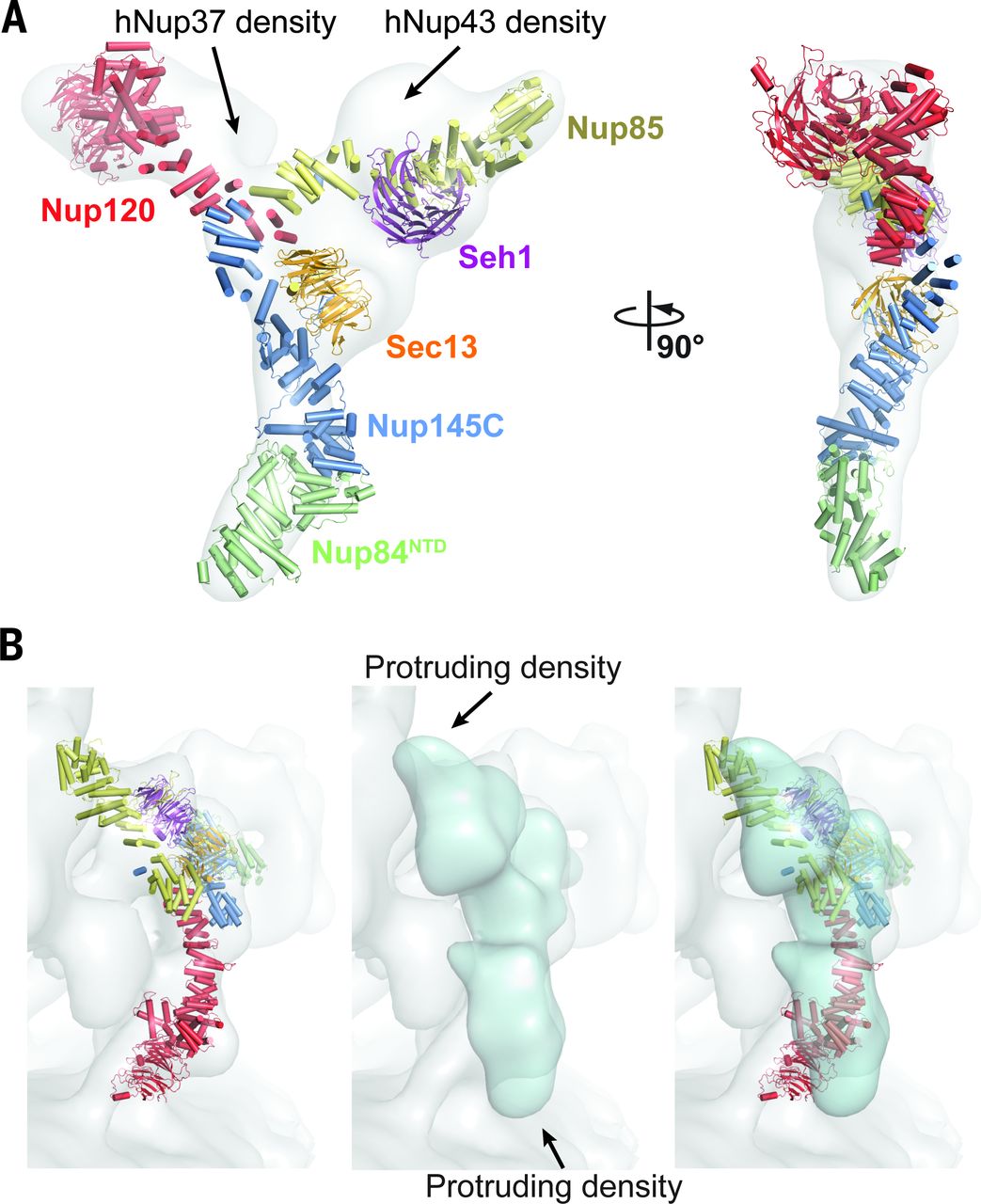
This will directly interface with efforts to crystallize or functionally trap some of these membrane proteins through conformationally-sensitive synthetic binders as chaperones and for functional/cell biological studies. Crystallization chaperones are a powerful way to increase the ability of a protein to crystallize. Crystallization chaperones are designed to bind to a protein, forming a complex that has new surfaces suitable for forming crystal contacts. The Koide and Kossiakoff labs, together with Sachdev S. Sidhu at Genentech, have developed a pipeline for producing SABs using phage display technology and a reduced complexity library of low complexity. These SABs are sensitive to the conformational state of the binder protein, and recent modifications have made it possible to adapt this technology to membrane protein binders (its most relevant example is the new crystal structure for full-length KcsA in complex with an SAB). The plan is to have individual targets in each of the major functional classes go through the selection under conditions that stabilize a variety of conformations. Identified SAB binders will be used for functional perturbation co-crystallization and eventual structural determinations.


