Summary of Progress
A. Specific Aims
Aim 1: To identify and characterize computationally and experimentally mutations and substrate/ligand settings that alter the conformational equilibrium of LeuT (between the states allowing access from the extracellular and intracellular sides, respectively). The combination of approaches will be applied iteratively to simulate and measure transport, binding, and dynamics (e.g., with EPR measurements of accessibility of engineered cysteines). This will establish experimental conditions that enrich differentially conformational states that we have already identified in the transport cycle.
Aim 2: To characterize the transitions between states and identify large scale rearrangements associated with the distinct conformational states established in Aim 1, using both continuous wave (CW) and double electron-electron resonance (DEER) EPR spectroscopy to measure spin-spin distances.
B. Studies and Results
Bridge 3, which now completed its first year of activity, comprises three synergistic components combining experimental and computational approaches in a collaboration between the Javitch, Mchaourab, and Weinstein labs. The Javitch lab has focused on functional characterization of LeuT, which, in conjunction with computational studies in the Weinstein lab, has uncovered a novel mechanism that depends upon the functional coupling of two substrate sites, the S1 and S2 sites, the latter of which is masked in the available crystal structures. Supported by another mechanism, the lab is also pursuing single molecule FRET (smFRET) experiments to probe the dynamics of LeuT in parallel to the computational simulations. This research is enhancing the scope of investigation and leverages the Glue grant context to disseminate smFRET data and mechanistic interpretations among the Bridge 3 collaborators and the Cores (see below). One example is the unprecedented opportunity that this Bridge provides to compare EPR-based findings with those from smFRET studies (see Abstracts below). The Mchaourab laboratory has focused within this first year of funding on the dynamic information obtained from a series of pairs of spin labels (48 pairs) strategically placed on both the extracellular and intracellular sides of LeuT to capture the expected opening and closing movements of these regions that must occur during the transport cycle. Together with the computational analysis methods, the approaches can be calibrated and interpreted with greater accuracy so as to advance our insights into the dynamics of this model system for secondary transport with an unprecedented combination of tools. The Javitch lab has also identified substrates, inhibitors, and mutations that bias the transporter conformation. Parallel computational studies in the Weinstein lab using various forms of biased and unbiased MD simulations have led to mechanistic insights that can be further evaluated using collaborative experiments with both smFRET and EPR. For example, the EPR studies in the Mchaourab lab have enabled us to identify and quantify conformational changes that are associated with various functional states of the transporter in a manner consonant with the computational simulations. The functional characterization and the computational simulations are applied together to evaluate the results from the EPR studies in an iterative manner that takes advantage of our findings regarding the ability of the Na+ ions and the transported substrates Leu and Ala, and of specific mutations, to bias the conformation of the transporter toward various functional states. To this end, the Mchaourab lab has made great progress this year in establishing robust methods for studying LeuT by DEER both in detergent and reconstituted into a lipid environment in nanodiscs (see Highlights below).
A major issue that we addressed this year was the controversy regarding the substrate binding and stoichiometry of LeuT. We discovered that binding to the S2 site can be elusive because LeuT is very sensitive to elevated detergent concentrations so that common protein purification methods can obscure the S2 site and thus impair the functional mechanism that drives transport (see Highlights below). This work has been accepted for publication in Nature Structural and Molecular Biology.
C. Collaborations
The Bridge has been enhanced by collaborations with the Computational Modeling Core through the work of the Weinstein lab, by Core 3 – the Synthetic Antigen Binder Core, in the form of collaboration between the Javitch and Kossiakoff labs, and by Core 1 – the Protein Expression Core, in the form of collaboration between the Mchaourab and Dotsch laboratory to achieve cell-free expression of LeuT. We have contributed new methods for the quantitative study of membrane protein interactions, and have screened for Fabs that bind LeuT in particular conformations and identified a panel of binders that will be characterized during the coming year. The plan is to use them to stabilize transporter conformations for the EPR studies in Bridge 3, as well as for crystallography and smFRET studies in the Javitch lab. Cell-free expression of LeuT directly into micelles or nanodiscs will circumvent many of the pitfalls and potential purification artifacts and simplify the production of mutants for EPR analysis.
D. Plans
In the coming year we plan to identify conformational changes at the intracellular and extracellular sides of LeuT that are associated with transport by completing an extensive series of DEER studies. These studies will continue to be guided by SMD and MD studies that are continuing in the Weinstein lab, as well as structural hypotheses that results from analyses of the resulting trajectories. In the coming year, the parallel computational studies aim to achieve a mechanistic description of the conformational transitions in LeuT in different environments (e.g. lipid bilayers and micelles of different lipid/detergent compositions) and to elucidate structural and functional determinants that determine the role of the membrane in the function and dynamics of LeuT. Special attention will be given to the generalizable insights about specific contributions from various membrane components and the effect of membrane remodeling on the functional mechanisms. The design of specific LeuT/detergent/lipid ratios in the planned simulations, and the probing of the mechanistic hypotheses pursued in the computational modeling and simulations, are enabled by the ongoing experimental studies in the Javitch and Mchaourab labs. The methodology for quantifying the membrane deformation by protein-membrane interactions, and the effects on membrane protein function are calculated with methods developed in the Weinstein lab for dissemination through the Computational Modeling Core, as they represent general approaches for the computational study of membrane protein function in the context of the cell membrane.
Thus, the activity of the consortium has already created an active context for exploration of new methods as well as the application of the technology and insights we have developed within this Bridge to the study of various membrane proteins. An additional example is the collaboration between the Mchaourab lab and Dr. Dötsch that has been developing approaches for in vitro translation of LeuT to expedite our ability to screen many mutants without laborious expression and purification (see Highlights below). The Mchaourab lab in collaboration with Dr, Glaubitz is also attempting to use solid state NMR to examine directly the binding of Leu to the S2 site (See Highlights below). The Mchaourab lab has also carried out preliminary studies with ApcT, a transporter from the amino acid, polyamine, and organocation (APC) transporter family that has a LeuT-like structural fold, in an attempt to expand our findings to other transporter families.
Research Highlights
Click to find out more about each research highlight (RH).
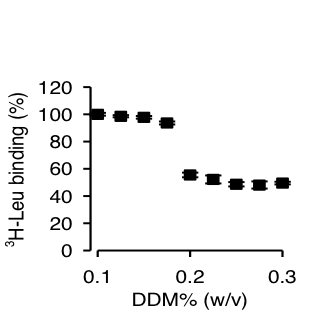
Figure 1. Effect of DDM on Leu binding by LeuT-WT.
Click to enlarge.High-resolution structures of LeuT, a bacterial NSS homologue, have provided a structural context for the exploration of transport mechanisms, but not a mechanistic understanding of the molecular events underlying transport. From results of computational studies using the crystal structures, and of binding and flux experiments, we have discovered a novel molecular mechanism of Na+-coupled symport that depends upon the binding of substrate in a second high affinity (S2) site. We show that substrate binding to the S2 site can be blocked during preparation of detergent-solubilized LeuT for functional studies and crystallization, thereby obscuring the functional mechanism of Na+-coupled symport. This brings to light the need to delineate the properties and mechanistic features of the allosteric membrane protein systems with appropriate attention to the environment in which they are studied.
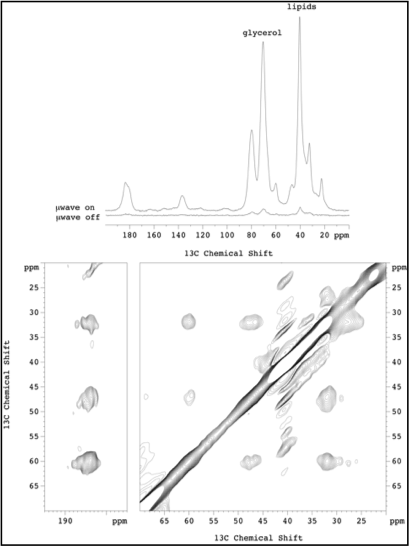
Figure 2.
Click to enlarge.A new opportunity to test and refine the molecular models developed in this Bridge Project, using specialized experimental approaches is being pursued in a collaboration initiated by the Mchaourab laboratory with Clemens Glaubitz at the University of Frankfurt. The aim is to use solid state NMR to probe the binding positions of leucine by detecting leucine bound at the S2 site and mapping the side chains to which it is coordinated. For this purpose, LeuT was uniformly labeled with 15N, reconstituted into liposomes in the presence of 13C leucine. Magic angle spinning solid state NMR analysis at 850 MHz was not sensitive enough to detect bound leucine. However, dynamic nuclear polarization at 393 MHz provided a 25-fold improvement in signal to noise (see upper panel in the Figure). The sensitivity improvement enabled a range of experiments to be carried out culminating in the detection of bound 13C Leu. 13C-13C correlation experiments (proton-driven spin diffusion) show intramolecular correlation between 13C within Leu. Different populations are observed, probably arising from bound and non-bound Leu. Experiments are in progress in samples without free leucine and using 15N-13C correlation to detect and assign the arginine side chain that coordinates leucine binding to S2.
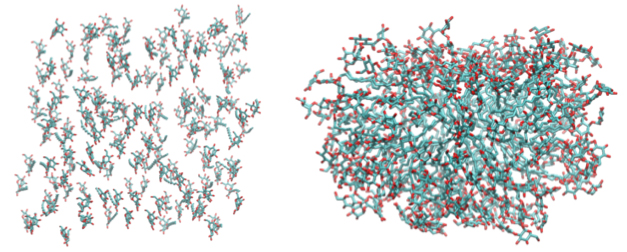
Figure 3: Configuration of 150 DDM molecules.
Click to enlarge.We have already taken some important initial steps towards the design and testing with MD simulations of LeuT in different detergent conditions, in comparison to its behavior in a lipid bilayer, that impact the functional properties of the transporter, as described above. In particular, to confirm whether available molecular force-fields for DDM (all-atom empirical potentials based on the most recent CHARMM hydrocarbon force-fields can reproduce spontaneous formation of DDM micelles as expected from experiment, we have simulated a self-assembly of DDM molecules into a micellar aggregate in solution from an unbiased all-atom MD simulations. Figure 3 shows initial and final (after 40 ns) configurations of the DDM molecules (water molecules removed for clarity), and illustrates the ability of 150 DDM molecules, a number consistent with reported aggregation number of 110-140 for DDM from the experimental studies, to spontaneously aggregate into a stable micelle.
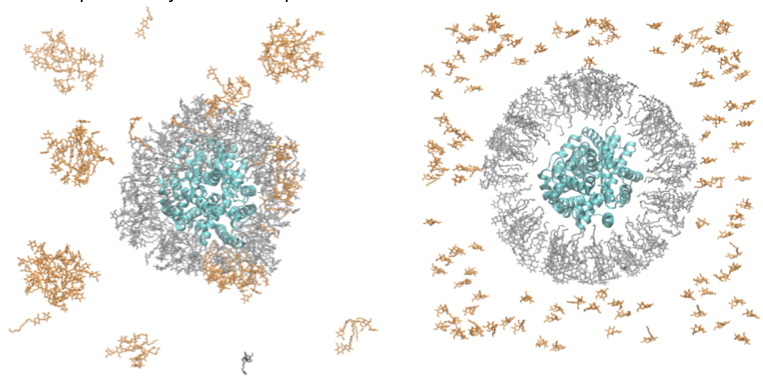
Figure 4: Snapshots from MD simulations of LeuT (PDB 3GJD) in DDM micelle.
Click to enlarge.Figure 4 shows a starting and a current (after 30 ns) snapshot of LeuT initially surrounded by a micelle containing 160 DDM molecules (Fig. 4A). The complete simulation box consisted of a single LeuT in complex with leucine ligand in the S1 site, and 2 Na+ and 1 Cl- ions (PDBid 3GJD), surrounded by 275 DDM molecules, 156814 water molecules, and additional 224 Cl- and 217 Na+ ions for electroneutrality (501441 total atoms). As can been seen from Fig. 4B, after 30ns of molecular dynamics, several important changes have occurred, which must be analyzed further: i) DDM molecules in the initial micelle have established tight packing around the LeuT; ii) monomeric DDMs have formed separate micellar structures; iii) some of the monomeric DDM molecules have merged into the center micelle, while one DDM from the original micelle has diffused away.
This preliminary simulation provides first clues about the environment surrounding LeuT in detergent. In order to arrive at equilibrium conditions for transporter/micelle complexes, it is essential to pursue the current simulation for much longer times and design a number of comparative/alternative setups for additional extensive simulations, and these are planned in the coming year.
LeuT has been crystallized bound with various amino acids (leucine, glycine, methionine, tyrosine and tryptophan), tricyclic antidepressants (TCA), and selective serotonin re-uptake inhibitors (SSRIs). Although these structures reveal molecular details of ion/substrate binding and inhibition, the conformational dynamics underlying a complete transport cycle have yet to be elucidated. Computational modeling studies have led to a proposal of a rocking-bundle mechanism by Forrest et al. that is based on the inherent inverted pseudo-symmetry between TM1-5 and TM6-10. According to this model, TM1, 2, 6, and 7 form a bundle that tilts with respect to other surrounding helices and results in alternate closing and opening of cytoplasmic and extracellular molecular gates. We are investigating the conformational changes of transmembrane helices predicted to move at different stages of the transport cycle by measuring distances between spin labels. Monitoring these dynamics should provide insights into molecular interactions that govern the extracellular gating of the transporter during the transport process. Our results show that the bundle does not behave as a single rigid body structural unit as previously proposed. Some TM helices within the bundle show a distinct increase in conformational flexibility as observed from the width of distance distributions, whereas others show a change in the equilibrium between different conformational intermediates in response to ion and or substrate binding. We are also analyzing these conformational changes with respect to the crystal structure through geometric optimization in order to consolidate distance changes into a model of the sodium bound intermediate.
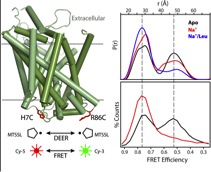
Figure 5.
Click to enlarge.This work will describe a systematic investigation of the intracellular dynamics of LeuT as well as detail progress in defining the LeuT transport mechanism and open-inward structure. Using site-directed spin labeling and double electron-electron resonance (DEER) spectroscopy, a pulsed EPR technique, over 40 double mutants were analyzed for their distance distributions and relative dynamics. Mechanistic descriptions were inferred by tracking shifts in equilibria between multi-component distance distributions and relating these shifts to biochemical conditions across all mutants. Closed-inward distances were largely consistent with the LeuT crystal structures, while primary gating motions were identified at helices 1 and 6 with additional reorientations of helices 5 and 7, resulting in the open-inward structure. Distance data are being implemented into restrained computational modeling to generate preliminary models of the open-inward conformation of LeuT. These results will be compared to the current models of LeuT open-in structure and transport mechanism derived using alternative approaches.
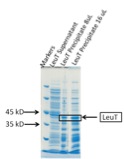
Figure 6.
Click to enlarge.We have used the protein core optimized cell-free expression system to express the integral membrane protein LeuT. Magnesium screens were performed to optimize the yield. However, we could not obtain sufficient amount of material for further detailed biochemical studies. A crucial parameter that influences the expression is the sequence at the N-terminus of the protein. Varying this sequence through adding different tags often results in significant increases in the expression yield. In the case of LeuT the originally used construct contained an N-terminal 10 His tag. A second construct that we used contained the His tag at the C-terminus and provided a higher basal expression level; we plan to continue to optimize the yield, first by performing a magnesium screen.
Mindful of the need to generalize inferences about specific membrane proteins in order to identify general principles in the context of Glue Grant goals, we are undertaking a study of ApcT, a member of the amino acid, polyamine, and organocation transporter family, which is a proton coupled amino acid transporter and has been proposed recently to share not only structural similarity with LeuT, but also key elements of its functional mechanism. Thus, ApcT consists of twelve transmembrane helices of which the first ten helices are a conserved 5+5 inverted repeat, similar to that of LeuT and AdiC. The prevalence of this conserved protein fold in ion coupled tranporters leads to the hypothesis of a conserved conformational mechanism for this family of transporters, including neurotransmitters transporters, as elaborated recently by members of the Consortium (Shi & Weinstein, 2010). The structure of ApcT was determined in the apo state by the Gouaux laboratory. Based on structural and sequence alignment with AdiC and LeuT, Shaffer et al. (2009) propose that TMH1 and TMH8 are involved in substrate binding. Protonation and deprotonation of K158 is important in transport activity, likening protonation and deprotonation to binding and release of a Na ion in LeuT.
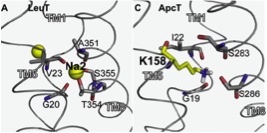
Figure 7.
Click to enlarge.To test the hypothesis of fold and mechanism conservation, we are investigating the rearrangements of helices during ion/substrate binding and release using spin labeling and electron paramagnetic resonance (EPR) spectroscopy. We have received the DNA construct of ApcT as a fusion protein to GFP from the Gouaux lab. We are in the process of removing the GFP tag and introducing unique pairs of cysteines to enable distance measurements between spin labels.