Efficient Exploration of Membrane-Associated Phenomena at Atomic Resolution
By Josh V. Vermaas, Javier L. Baylon, Mark J. Arcario, Melanie P. Muller, Zhe Wu, Taras V. Pogorelov, and Emad Tajkhorshid.
Published in J Membr Biol. 2015 Jun;248(3):563-82.
PMID: 25998378. PMCID: PMC4490090. Link to publication page.
Core: Computational Modeling Core.
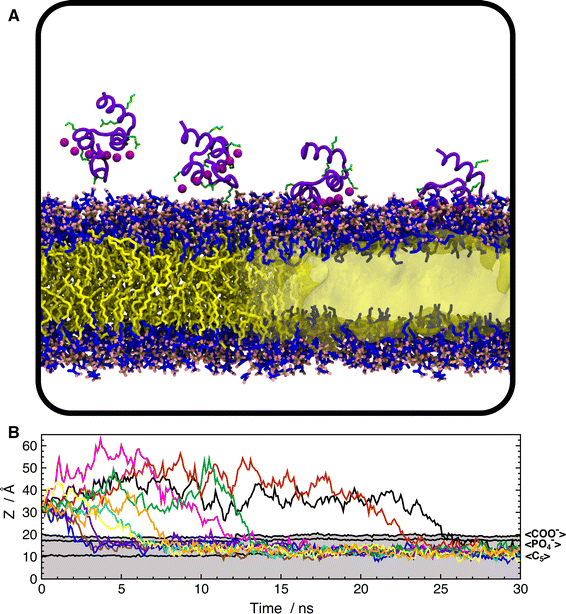
Figure 1. Representative membrane transporters and functional events studied recently. Proteins are shown in cartoon representation, with the lipid bilayer in the background. Each transporter respectively represents a well-documented superfamily/family of membrane transporters in recent computational studies. Four close-up views of local conformations relevant with several critical functional events, that is, Na+-binding (left), H+-binding (middle left), ATP binding (middle right), and substrate binding (right).
Abstract
Biological membranes constitute a critical component in all living cells. In addition to providing a conducive environment to a wide range of cellular processes, including transport and signaling, mounting evidence has established active participation of specific lipids in modulating membrane protein function through various mechanisms. Understanding lipid–protein interactions underlying these mechanisms at a sufficiently high resolution has proven extremely challenging, partly due to the semi-fluid nature of the membrane. In order to address this challenge computationally, multiple methods have been developed, including an alternative membrane representation termed highly mobile membrane mimetic (HMMM) in which lateral lipid diffusion has been significantly enhanced without compromising atomic details. The model allows for efficient sampling of lipid–protein interactions at atomic resolution, thereby significantly enhancing the effectiveness of molecular dynamics simulations in capturing membrane-associated phenomena. In this review, after providing an overview of HMMM model development, we will describe briefly successful application of the model to study a variety of membrane processes, including lipid-dependent binding and insertion of peripheral proteins, the mechanism of phospholipid insertion into lipid bilayers, and characterization of optimal tilt angle of transmembrane helices. We conclude with practical recommendations for proper usage of the model in simulation studies of membrane processes.