Expression, purification, and reconstitution of the voltage-sensing domain from Ci-VSP
By Qufei Li, Vishwanath Jogini, Sherry Wanderling, D. Marien Cortes, and Eduardo Perozo.
Published in Biochemistry 51(41): 8132-8142 (2012) on September 18, 2012. PMID: 22989304. Link to Pubmed page.
Core Facility: Membrane Protein Expression/Purification
Abstract
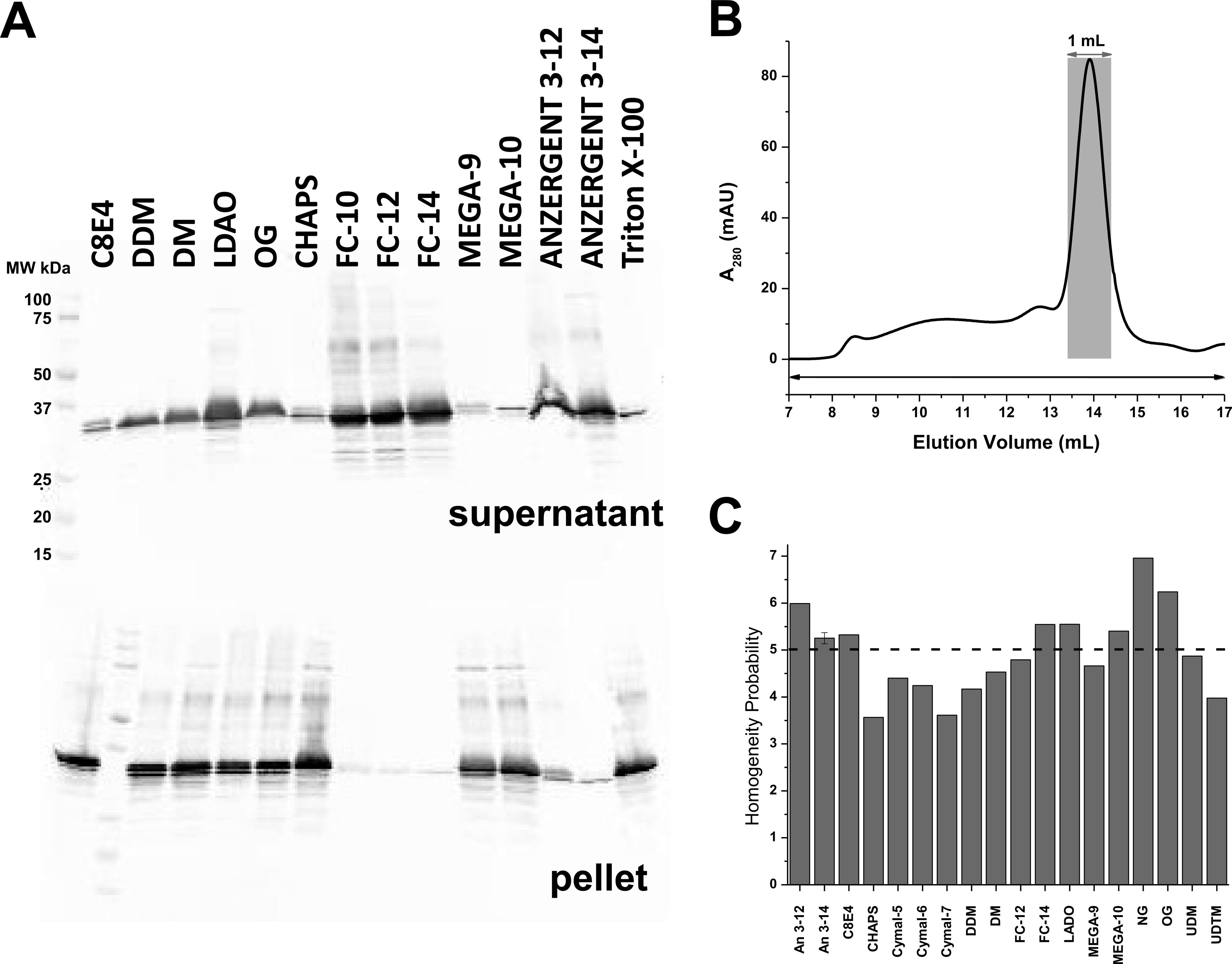
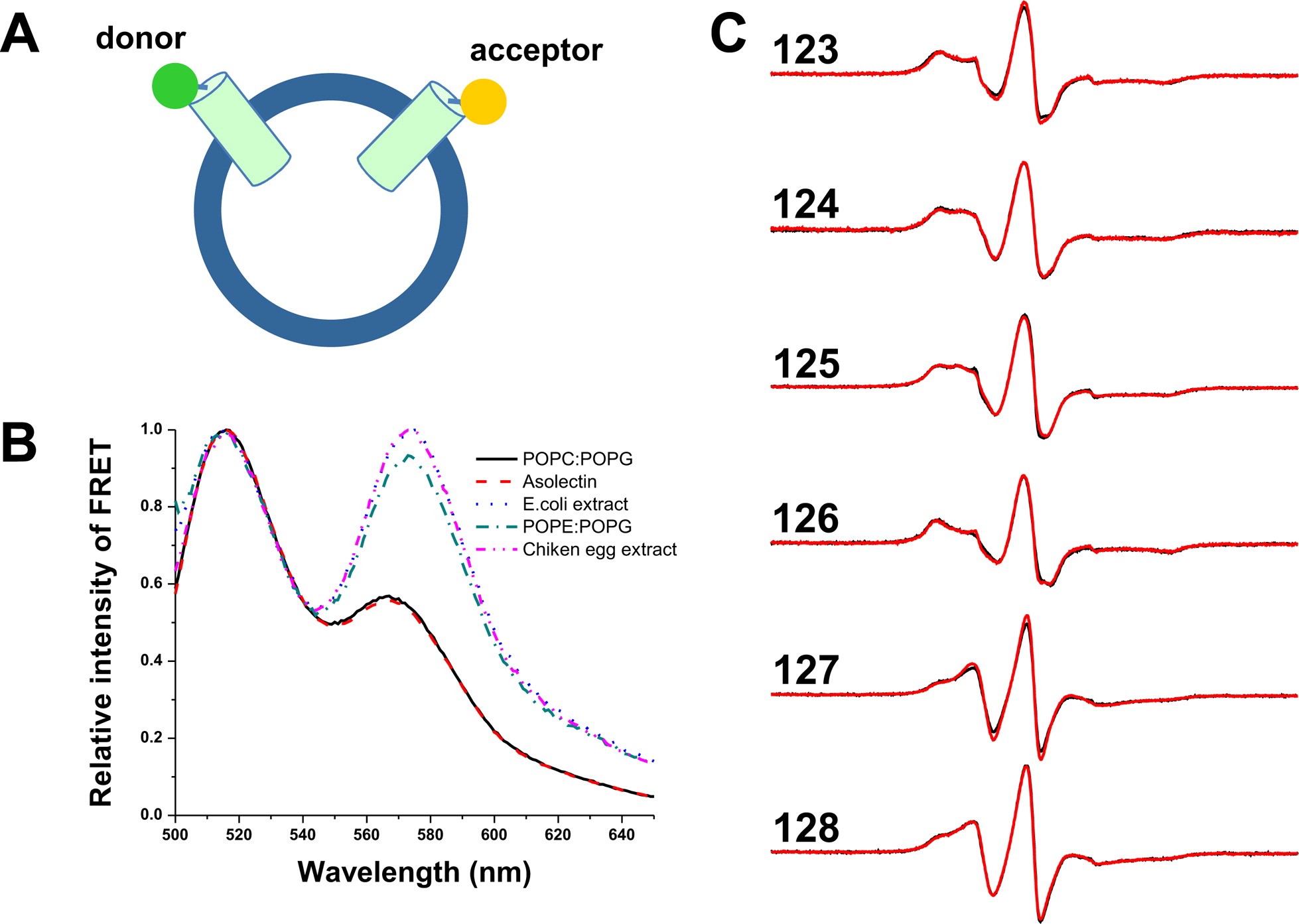
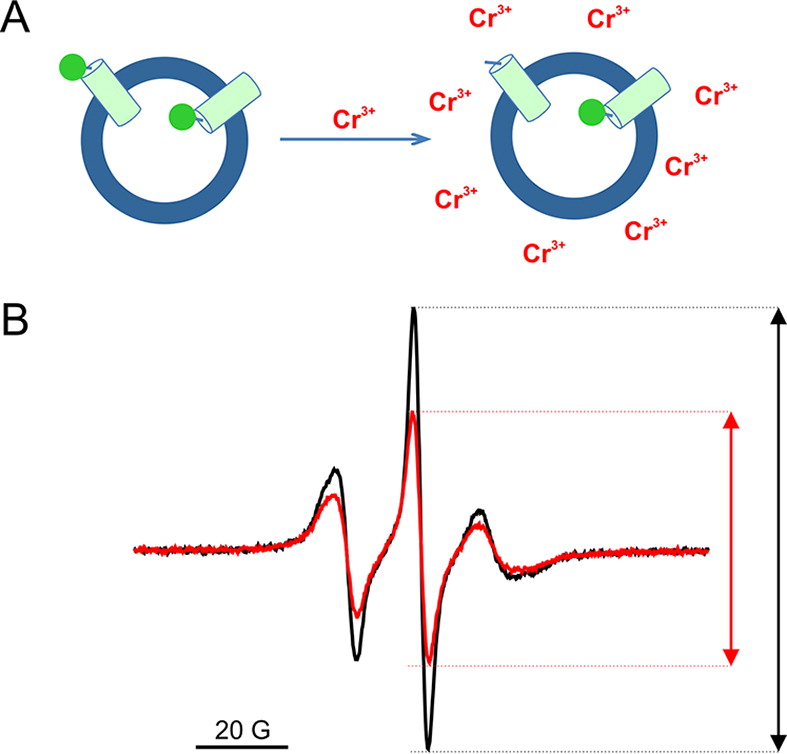
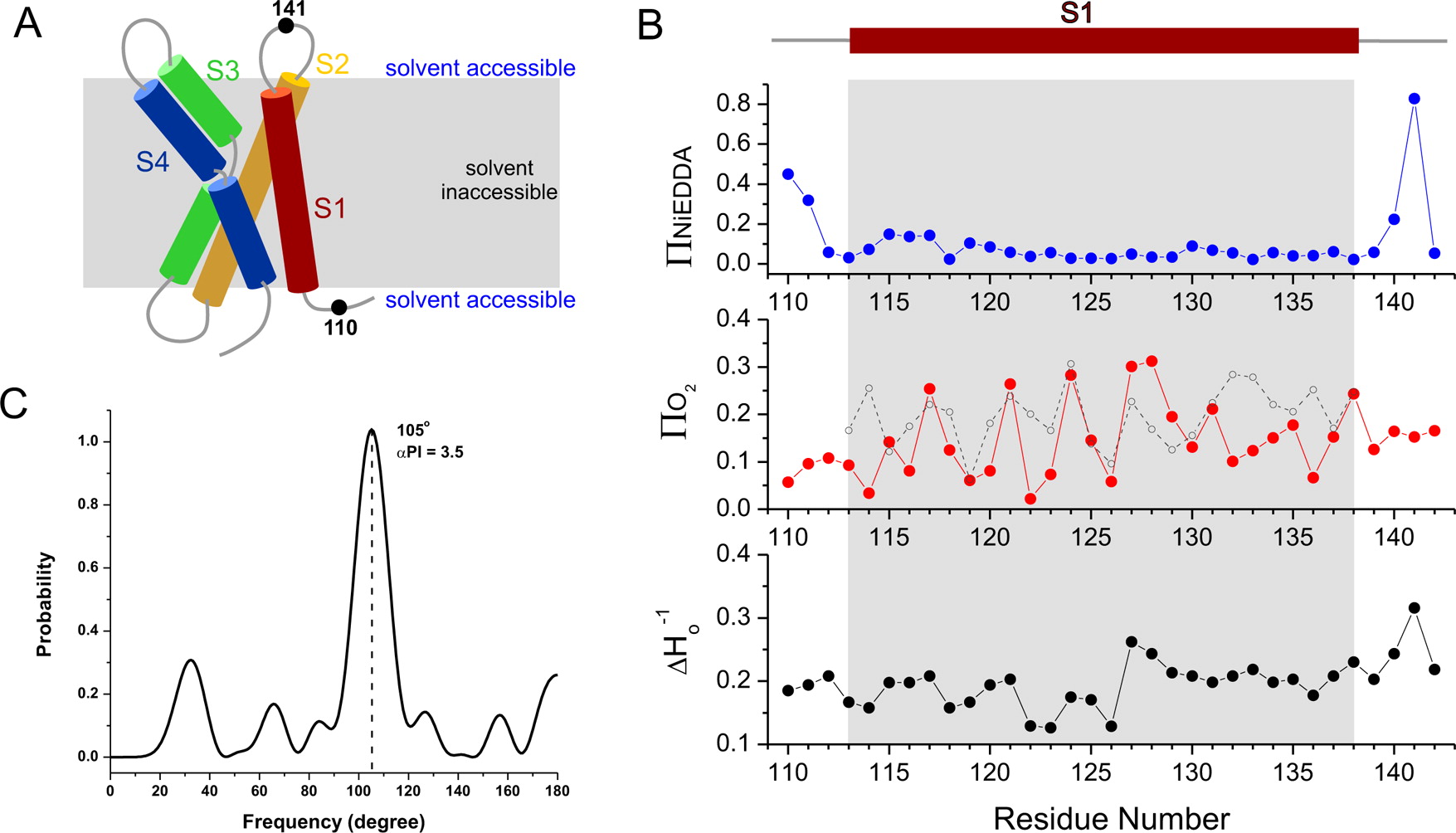
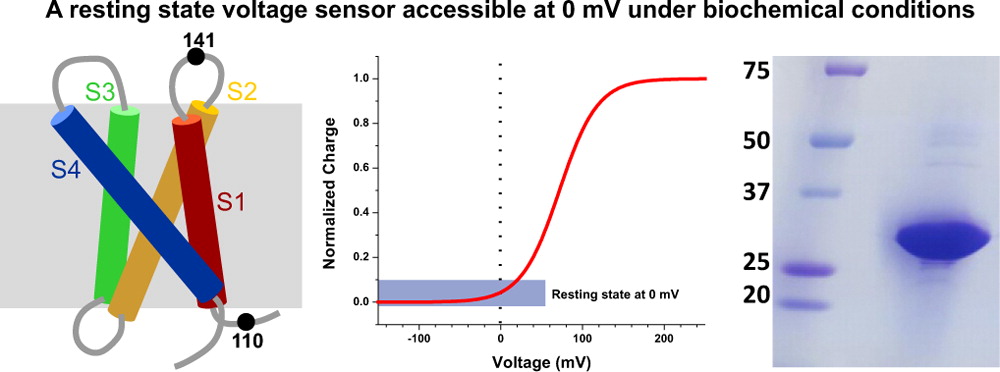
The voltage-sensing domain (VSD) is the common scaffold responsible for the functional behavior of voltage-gated ion channels, voltage sensitive enzymes, and proton channels. Because of the position of the voltage dependence of the available VSD structures, at present, they all represent the activated state of the sensor. Yet in the absence of a consensus resting state structure, the mechanistic details of voltage sensing remain controversial. The voltage dependence of the VSD from Ci-VSP (Ci-VSD) is dramatically right shifted, so that at 0 mV it presumably populates the putative resting state. Appropriate biochemical methods are an essential prerequisite for generating sufficient amounts of Ci-VSD protein for high-resolution structural studies. Here, we present a simple and robust protocol for the expression of eukaryotic Ci-VSD in Escherichia coli at milligram levels. The protein is pure, homogeneous, monodisperse, and well-folded after solubilization in Anzergent 3-14 at the analyzed concentration ( 0.3 mg/mL). Ci-VSD can be reconstituted into liposomes of various compositions, and initial site-directed spin labeling and electron paramagnetic resonance (EPR) spectroscopic measurements indicate its first transmembrane segment folds into an α-helix, in agreement with the homologous region of other VSDs. On the basis of our results and enhanced relaxation EPR spectroscopy measurement, Ci-VSD reconstitutes essentially randomly in proteoliposomes, precluding straightforward application of transmembrane voltages in combination with spectroscopic methods. Nevertheless, these results represent an initial step that makes the resting state of a VSD accessible to a variety of biophysical and structural approaches, including X-ray crystallography, spectroscopic methods, and electrophysiology in lipid bilayers.
0.3 mg/mL). Ci-VSD can be reconstituted into liposomes of various compositions, and initial site-directed spin labeling and electron paramagnetic resonance (EPR) spectroscopic measurements indicate its first transmembrane segment folds into an α-helix, in agreement with the homologous region of other VSDs. On the basis of our results and enhanced relaxation EPR spectroscopy measurement, Ci-VSD reconstitutes essentially randomly in proteoliposomes, precluding straightforward application of transmembrane voltages in combination with spectroscopic methods. Nevertheless, these results represent an initial step that makes the resting state of a VSD accessible to a variety of biophysical and structural approaches, including X-ray crystallography, spectroscopic methods, and electrophysiology in lipid bilayers.