Molecular dynamics investigation of the ω-current in the Kv1.2 voltage sensor domains
By Fatemeh Khalili-Araghi, Emad Tajkhorshid, Benoît Roux, and Klaus Schulten.
Published in Biophysical Journal 102(2): 258-267 on January 17, 2012.
PMID: 22339862. PMCID: PMC3260662. Link to Pubmed page.
Core Facility: Computational Modeling
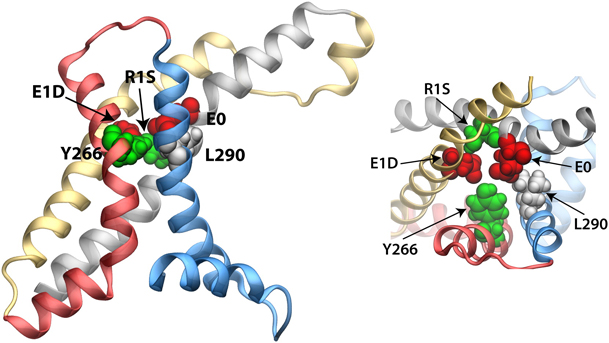
Figure 7. Maximum constriction region near the center of the VSD (−1 Å < z < 6 Å). Residues E0, E1D, R1S, L290, and Y266 are highlighted (van der Waals representation) and colored according to their chemical properties, namely, acidic (red), basic (blue), hydrophobic (white), and hydrophilic (green).
Abstract
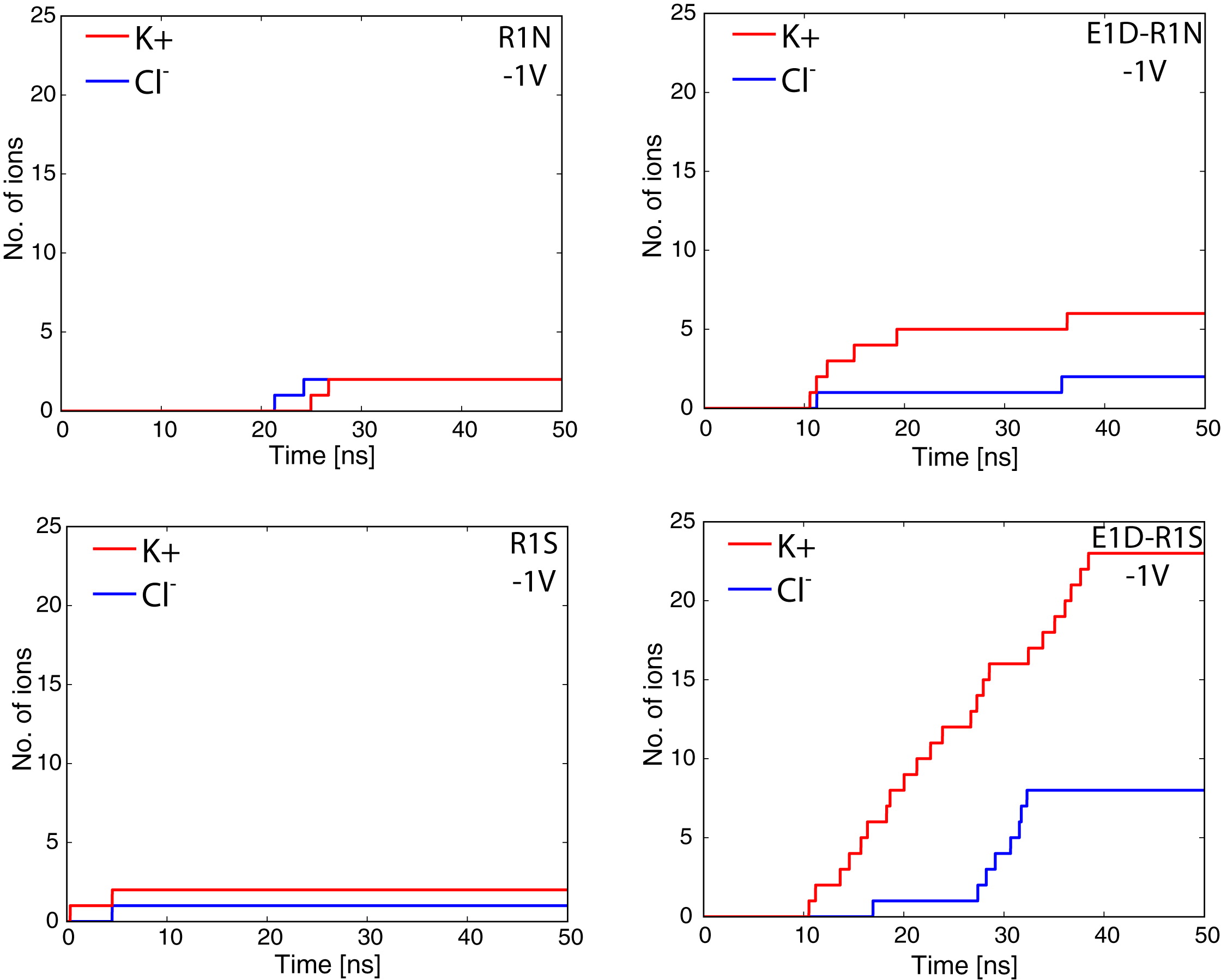
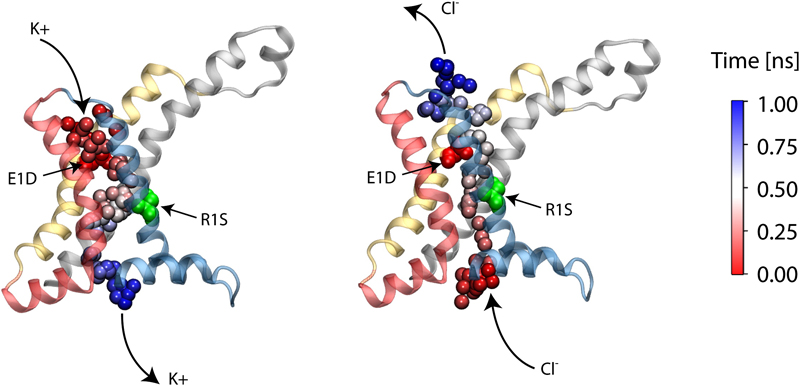
Voltage sensor domains (VSD) are transmembrane proteins that respond to changes in membrane voltage and modulate the activity of ion channels, enzymes, or in the case of proton channels allow permeation of protons across the cell membrane. VSDs consist of four transmembrane segments, S1–S4, forming an antiparallel helical bundle. The S4 segment contains several positively charged residues, mainly arginines, located at every third position along the helix. In the voltage-gated Shaker K+ channel, the mutation of the first arginine of S4 to a smaller uncharged amino acid allows permeation of cations through the VSD. These currents, known as ω-currents, pass through the VSD and are distinct from K+ currents passing through the main ion conduction pore. Here we report molecular dynamics simulations of the ω-current in the resting-state conformation for Kv1.2 and for four of its mutants. The four tested mutants exhibit various degrees of conductivity for K+ and Cl- ions, with a slight selectivity for K+ over Cl-. Analysis of the ion permeation pathway, in the case of a highly conductive mutant, reveals a negatively charged constriction region near the center of the membrane that might act as a selectivity filter to prevent permeation of anions through the pore. The residues R1 in S4 and E1 in S2 are located at the narrowest region of the ω-pore for the resting state conformation of the VSD, in agreement with experiments showing that the largest increase in current is produced by the double mutation E1D and R1S.