Nano-Positioning System for Structural Analysis of Functional Homomeric Proteins in Multiple Conformations
By H. Clark Hyde, Walter Sandtner, Ernesto Vargas, Alper T. Dagcan, Janice L. Robertson, Benoît Roux, Ana M. Correa, and Francisco Bezanilla.
Published in Structure 20(10): 1629-1640 (2012) on October 10, 2012. PMID: 23063010. PMCID: PMC3283816. Link to Pubmed page.
Project: Conformational Transitions in P-class ATPases. Core Facility: Computational Modeling.
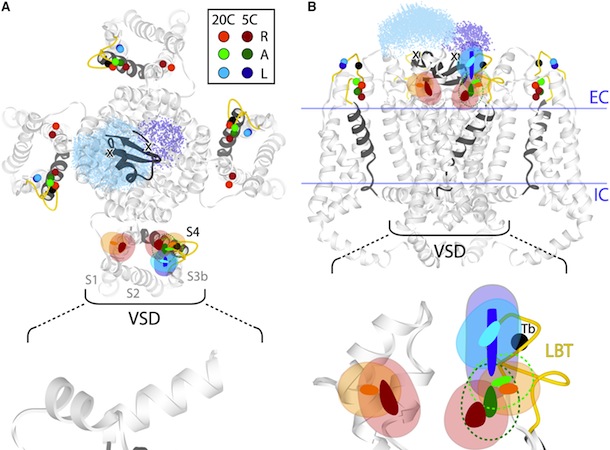
Figure 5. Structural Rearrangements of Shaker S4 Determined by SNPS. (A and B) Donor positions were determined by SNPS with diffusion modeling for the Shaker S4(4) LBT construct measured in the resting ‘‘R’’, active ‘‘A’’, and relaxed ‘‘L’’ functional states (see key for color scheme). Displayed solution types are resting (1 and 2), active (2), and relaxed (2). Mapped donor positions are shown for the S4(4) LBT construct measured using two different toxin labeling sites: AgTx2-D20C and AgTx2-N5C, superimposed on our relaxed state S4(4) LBT structural model with LBT (yellow) and encaged terbium(black). AgTx2(II) is shown docked to the pore with both calibrated acceptor clouds: D20C (cyan dots) and N5C (deep blue dots); cysteine Ca positions are labeled ‘‘X’’. Mean donor positions are represented as spheres. In one subunit, 95% confidence surfaces are shown for the mean position (solid inner surface) and diffusion cloud (transparent outer surface) of each donor. Viewing angles are extracellular in (A) and side in (B). The hydrophobic thickness of Kv1.2 is depicted by extracellular (EC) and intracellular (IC) lines, obtained from OPM (PDB 3LUT). Bottom: magnified views of one subunit’s VSD; scale bar is 5 Å. See also Figures S7 and S8.
Abstract
Proteins may undergo multiple conformational changes required for their function. One strategy used to estimate target-site positions in unknown structural conformations involves single-pair resonance energy transfer (RET) distancemeasurements. However, interpretation of inter-residue distances is difficult when applied to three-dimensional structural rearrangements, especially in homomeric systems. We developed a positioning method using inverse trilateration/triangulation to map target sites within a homomeric protein in all defined states, with simultaneous functional recordings. The procedure accounts for probe diffusion to accurately determine the three-dimensional position and confidence region of lanthanide LRET donors attached to a target site (one per subunit), relative to a single fluorescent acceptor placed in a static site. As first application, the method is used to determine the position of a functional voltage-gated potassium channel’s voltage sensor. Our results verify the crystal structure relaxed conformation and report on the resting and active conformations for which crystal structures are not available.