Navigating Membrane Protein Structure, Dynamics, and Energy Landscapes Using Spin Labeling and EPR Spectroscopy
By Derek P. Claxton, Kelli Kazmier, Smriti Mishra, and Hassane S Mchaourab.
Published in Methods Enzymol. Epub 2015 Aug 29. PMID: 26477257. PMCID: 5222538. Link to Pubmed page.
Core Facility: Spectroscopy and Instrumentation.
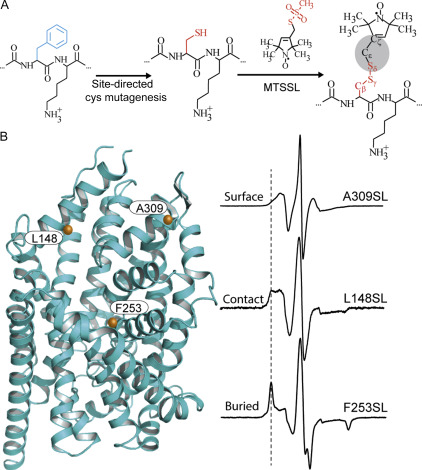
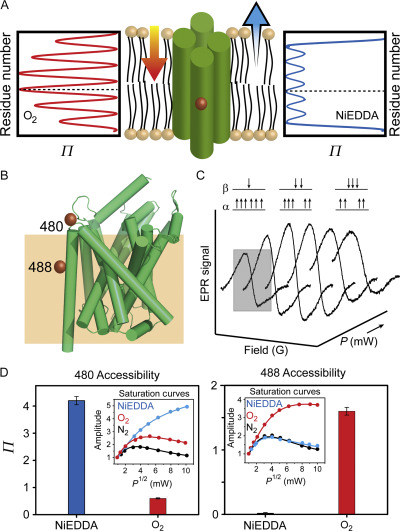
Figure 3. Site-directed spin labeling and correlation of the EPR spectrum with local structure. (A) Targeted cysteine mutagenesis introduces a sulfhydryl moiety for the attachment of a nitroxide spin label, such as MTSSL. Rotational isomerization of MTSSL predominantly around the bonds highlighted in gray is reflected in the EPR spectral lineshape. (B) The degree of rotational freedom of the label is determined by the local packing environment. Fast rotational correlation times (~ 1 ns) correspond to spin labels attached to surface-exposed sites. Tertiary contact interactions or buried sites that restrict spin label motion reduce the rate and amplitude of isomerization leading to broadening of the lineshape. The dashed line emphasizes the progressive appearance of a slow-motion component associated with restricted rotation.
Abstract
A detailed understanding of the functional mechanism of a protein entails the characterization of its energy landscape. Achieving this ambitious goal requires the integration of multiple approaches including determination of high-resolution crystal structures, uncovering conformational sampling under distinct biochemical conditions, characterizing the kinetics and thermodynamics of transitions between functional intermediates using spectroscopic techniques, and interpreting and harmonizing the data into novel computational models. With increasing sophistication in solution-based and ensemble-oriented biophysical approaches such as electron paramagnetic resonance (EPR) spectroscopy, atomic resolution structural information can be directly linked to conformational sampling in solution. Here, we detail how recent methodological and technological advances in EPR spectroscopy have contributed to the elucidation of membrane protein mechanisms. Furthermore, we aim to assist investigators interested in pursuing EPR studies by providing an introduction to the technique, a primer on experimental design, and a description of the practical considerations of the method toward generating high quality data.