Structural Refinement from Restrained-Ensemble Simulations Based on EPR/DEER Data: Application to T4 Lysozyme
By Shahidul Islam, Richard Stein, Hassane Mchaourab and Benoı̂t Roux.
Published in Journal of Physical Chemistry B [Epub ahead of print] March 19, 2013;117(17):4740-54. PMID: 23510103. PMCID: PMC3684008. Link to Pubmed page.
Project: Structural Dynamics of ABC Transporter. Core Facility: Computational Modeling
Abstract
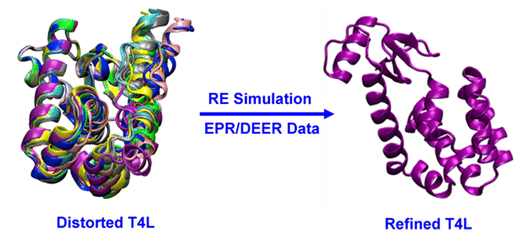
DEER (double electron–electron resonance) is a powerful pulsed ESR (electron spin resonance) technique allowing the determination of distance histograms between pairs of nitroxide spin-labels linked to a protein in a native-like solution environment. However, exploiting the huge amount of information provided by ESR/DEER histograms to refine structural models is extremely challenging. In this study, a restrained ensemble (RE) molecular dynamics (MD) simulation methodology is developed to address this issue. In RE simulation, the spin–spin distance distribution histograms calculated from a multiple-copy MD simulation are enforced, via a global ensemble-based energy restraint, to match those obtained from ESR/DEER experiments. The RE simulation is applied to 51 ESR/DEER distance histogram data from spin-labels inserted at 37 different positions in T4 lysozyme (T4L). The rotamer population distribution along the five dihedral angles connecting the nitroxide ring to the protein backbone is determined and shown to be consistent with available information from X-ray crystallography. For the purpose of structural refinement, the concept of a simplified nitroxide dummy spin-label is designed and parametrized on the basis of these all-atom RE simulations with explicit solvent. It is demonstrated that RE simulations with the dummy nitroxide spin-labels imposing the ESR/DEER experimental distance distribution data are able to systematically correct and refine a series of distorted T4L structures, while simple harmonic distance restraints are unsuccessful. This computationally efficient approach allows experimental restraints from DEER experiments to be incorporated into RE simulations for efficient structural refinement.


