A highly tilted membrane configuration for the prefusion state of synaptobrevin
By Andrew E. Blanchard, Mark J. Arcario, Klaus Schulten, and Emad Tajkhorshid.
Published in Biophys J. 2014 Nov 4;107(9):2112-21.
PMID: 25418096. PMCID: PMC4223215 [Available on 2015/11/4]. Link to publication page.
Core: Computational Modeling Core.
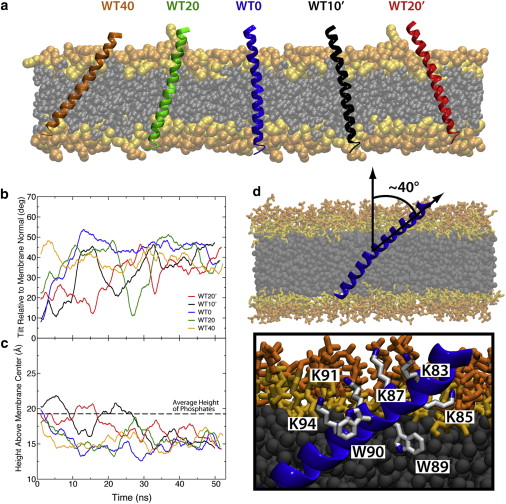
Figure 2. Membrane-embedded synaptobrevin. (a) Initial conditions for simulations with wild-type synaptobrevin varied in both initial insertion depth and tilt. The simulations are labeled according to starting tilt with respect to the membrane normal. Initial tilts with the lysines in the linker region (residues 87, 91, and 94) angled toward the center of the membrane are denoted with a prime (′). (b) Running average of the tilt angle between TM1 and the membrane normal. The tilt angle converges after ∼30 ns. (c) Running average of the height of the tryptophan residues above the midpoint of the membrane. The distance below the phosphate layer of the membrane (dashed line) for the tryptophan residues converges to a value of ∼5 Å. (d) The highly tilted state resulting in all cases is characterized by lysine residues aligned along one side of the synaptobrevin helix and tryptophan residues inserted into the membrane on the other side of the helix. To see this figure in color, go online.
Abstract
The SNARE complex plays a vital role in vesicle fusion arising during neuronal exocytosis. Key components in the regulation of SNARE complex formation, and ultimately fusion, are the transmembrane and linker regions of the vesicle-associated protein, synaptobrevin. However, the membrane-embedded structure of synaptobrevin in its prefusion state, which determines its interaction with other SNARE proteins during fusion, is largely unknown. This study reports all-atom molecular-dynamics simulations of the prefusion configuration of synaptobrevin in a lipid bilayer, aimed at characterizing the insertion depth and the orientation of the protein in the membrane, as well as the nature of the amino acids involved in determining these properties. By characterizing the structural properties of both wild-type and mutant synaptobrevin, the effects of C-terminal additions on tilt and insertion depth of membrane-embedded synaptobrevin are determined. The simulations suggest a robust, highly tilted state for membrane-embedded synaptobrevin with a helical connection between the transmembrane and linker regions, leading to an apparently new characterization of structural elements in prefusion synaptobrevin and providing a framework for interpreting past mutation experiments.