
Cartoon model of LeuT transport derived from EPR data, click to enlarge (Publication source)
Using the recently determined crystal structure of a prokaryotic leucine transporter (LeuT), the Transport Cycle in Neurotransmitter Uptake Project explores conformational changes and dynamic properties relevant to function in Neurotransmitter transporters translocation cycle using a combination of computational, functional, and spectroscopic approaches.
The conceptual design, scope and integration of this Project exemplifies the consortium approach to discovery of mechanistic principles of secondary active transport, including the conformational dynamics that govern alternating access in transporters and the allosteric interplay of substrate sites that transduce the energy stored in the electrochemical Na+ gradient into transport work. One of the most recent collaborative publications from members of this Project (lab of Harel Weinstein) and the Consortium at large (labs of Consortium member Olga Boudker and associate member Scott Blanchard) describes such conformational dynamics for a complex transporter system, illustrating the power of the combined experimental / computational approaches we have implemented for the study of these membrane proteins in the possible next phase of the grant:
Akyuz N, Georgieva ER, Zhou Z, Stolzenberg S, Cuendet MA, Khelashvili G, Altman RB, Terry DS, Freed JH, Weinstein H, Boudker O, Blanchard SC.
Transport domain unlocking sets the uptake rate of an aspartate transporter
Nature. 2015 Feb 5;518(7537):68-73. PMID: 25652997. PMCID: PMC4351760.
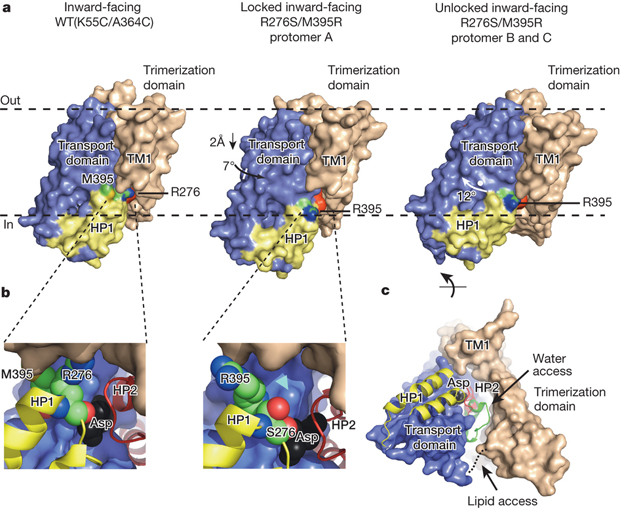
Crystal structure of the H276,395-GltPh.
Publication abstract: Glutamate transporters terminate neurotransmission by clearing synaptically released glutamate from the extracellular space, allowing repeated rounds of signalling and preventing glutamate-mediated excitotoxicity. Crystallographic studies of a glutamate transporter homologue from the archaeon Pyrococcus horikoshii, GltPh, showed that distinct transport domains translocate substrates into the cytoplasm by moving across the membrane within a central trimerization scaffold. Here we report direct observations of these ‘elevator-like’ transport domain motions in the context of reconstituted proteoliposomes and physiological ion gradients using single-molecule fluorescence resonance energy transfer (smFRET) imaging. We show that GltPh bearing two mutations introduced to impart characteristics of the human transporter exhibits markedly increased transport domain dynamics, which parallels an increased rate of substrate transport, thereby establishing a direct temporal relationship between transport domain motion and substrate uptake. Crystallographic and computational investigations corroborated these findings by revealing that the ‘humanizing’ mutations favour structurally ‘unlocked’ intermediate states in the transport cycle exhibiting increased solvent occupancy at the interface between the transport domain and the trimeric scaffold.
In a second publication from this Project, Matthias Quick (who is working with Project PI Jonathan Javitch), and Computational Modeling Core participant Lei Shi (from Harel Weinstein’s group), published findings that reveal the existence of two substrate sites in vSGLT, a member of the solute-sodium symporter (SSS) family of secondary transporters, as well as in PutP, another member of the SSS family. This publication follows a number of Consortium-sponsored interactions between Quick and Shi as well as Javitch and Weinstein. An earlier 2013 research initiative examined the aggregation dynamics and detergent binding to the second substrate site binding site of LeuT, and in the same year, their team published a key paper on the chloride binding site of neurotransmitter sodium transporters. The very first paper published in the name of the Project, was also the product of collaboration between this team of scientists, and showed how experimental conditions can block the second high-affinity site of LeuT. The important discovery of the present publication suggests that our findings in LeuT are generalizable to a different family of secondary transporters. These findings are the basis for forthcoming scientific collaborations between Quick and Shi, further demonstrating the broadening of the impact of the consortium and the synergy that the “glue” of the consortium has engendered.
Li Z, Lee AS, Bracher S, Jung H, Paz A, Kumar JP, Abramson J, Quick M, Shi L.
Identification of a second substrate-binding site in solute-sodium symporters
J Biol Chem., 2015 Jan 2;290(1):127-41. PMID: 25398883. PMCID: PMC4281715.
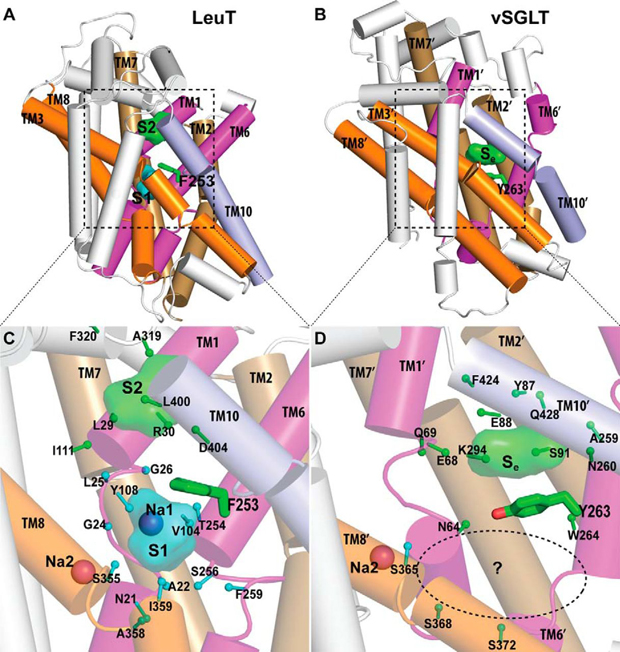
Crystallographically identified substrate-binding site of vSGLT is located more extracellularly than that of LeuT.
Publication abstract: The structure of the sodium/galactose transporter (vSGLT), a solute-sodium symporter (SSS) from Vibrio parahaemolyticus, shares a common structural fold with LeuT of the neurotransmitter-sodium symporter family. Structural alignments between LeuT and vSGLT reveal that the crystallographically identified galactose-binding site in vSGLT is located in a more extracellular location relative to the central substrate-binding site (S1) in LeuT. Our computational analyses suggest the existence of an additional galactose-binding site in vSGLT that aligns to the S1 site of LeuT. Radiolabeled galactose saturation binding experiments indicate that, like LeuT, vSGLT can simultaneously bind two substrate molecules under equilibrium conditions. Mutating key residues in the individual substrate-binding sites reduced the molar substrate-to-protein binding stoichiometry to ∼1. In addition, the related and more experimentally tractable SSS member PutP (the Na+/proline transporter) also exhibits a binding stoichiometry of 2. Targeting residues in the proposed sites with mutations results in the reduction of the binding stoichiometry and is accompanied by severely impaired translocation of proline. Our data suggest that substrate transport by SSS members requires both substrate-binding sites, thereby implying that SSSs and neurotransmitter-sodium symporters share common mechanistic elements in substrate transport.







